Tilman, D., Balzer, C., Hill, J. & Befort, B. L. Global food demand and the sustainable intensification of agriculture. Proc. Natl Acad. Sci. USA 108, 20260–20264 (2011).
Hunter, M. C., Smith, R. G., Schipanski, M. E., Atwood, L. W. & Mortensen, D. A. Agriculture in 2050: recalibrating targets for sustainable intensification. Bioscience 67, 386–391 (2017).
Cunningham, S. A. et al. To close the yield-gap while saving biodiversity will require multiple locally relevant strategies. Agric. Ecosyst. Environ. 173, 20–27 (2013).
Popp, J., Petö, K. & Nagy, J. Pesticide productivity and food security. A review. Agron. Sustain. Dev. 33, 243–255 (2013).
Oerke, E.-C. Crop losses to pests. J. Agric. Sci. 144, 31–43 (2006).
Singh, S. K., Hodda, M. & Ash, G. J. Plant-parasitic nematodes of potential phytosanitary importance, their main hosts and reported yield losses. EPPO Bull. 43, 334–374 (2013).
Abad, P. et al. Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nat. Biotechnol. 26, 909–915 (2008).
Jones, R. K. in Nematology in South Africa: A View from the 21st Century (eds Fourie, H. et al.) 129–150 (Springer, 2017).
Desaeger, J., Wram, C. & Zasada, I. New reduced-risk agricultural nematicides—rationale and review. J. Nematol. 52, e2020-911 (2020).
EU Pesticides Database, https://ec.europa.eu/food/plant/pesticides/eu-pesticides-database/start/screen/active-substances (European Commission).
NPIC Product Research Online (NPRO), http://npic.orst.edu/NPRO/# (National Pesticide Information Center, accessed 25 July 2022).
Jordan, S., Nischwitz, C., Ramirez, R. & Gordillo, L. F. Managing the spread of alfalfa stem nematodes (Ditylenchus dipsaci): the relationship between crop rotation periods and pest reemergence. Nat. Resour. Model. 30, e12083 (2017).
d’Errico, G., Giacometti, R., Roversi, P. F., d’Errico, F. P. & Woo, S. L. Mode of action and efficacy of iprodione against the root-knot nematode Meloidogyne incognita. Ann. Appl. Biol. 171, 506–510 (2017).
Burns, A. R. et al. Caenorhabditis elegans is a useful model for anthelmintic discovery. Nat. Commun. 6, 7485 (2015).
Culetto, E. et al. The Caenorhabditis elegans unc-63 gene encodes a levamisole-sensitive nicotinic acetylcholine receptor α subunit. J. Biol. Chem. 279, 42476–42483 (2004).
Hu, Y., Xiao, S. H. & Aroian, R. V. The new anthelmintic tribendimidine is an l-type (levamisole and pyrantel) nicotinic acetylcholine receptor agonist. PLoS Negl. Trop. Dis. 3, e499 (2009).
Driscoll, M., Dean, E., Reilly, E., Bergholz, E. & Chalfie, M. Genetic and molecular analysis of a Caenorhabditis elegans β-tubulin that conveys benzimidazole sensitivity. J. Cell Biol. 109, 2993–3003 (1989).
Dent, J. A., Smith, M. M., Vassilatis, D. K. & Avery, L. The genetics of ivermectin resistance in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 97, 2674–2679 (2000).
Kaminsky, R. et al. A new class of anthelmintics effective against drug-resistant nematodes. Nature 452, 176–180 (2008).
Guest, M. et al. The calcium-activated potassium channel, SLO-1, is required for the action of the novel cyclo-octadepsipeptide anthelmintic, emodepside, in Caenorhabditis elegans. Int. J. Parasitol. 37, 1577–1588 (2007).
Kawasaki, I., Jeong, M. H., Oh, B. K. & Shim, Y. H. Apigenin inhibits larval growth of Caenorhabditis elegans through DAF-16 activation. FEBS Lett. 584, 3587–3591 (2010).
Rand, J. B. & Russell, R. L. Choline acetyltransferase-deficient mutants of the nematode Caenorhabditis elegans. Genetics 106, 227–248 (1984).
Hu, Y., Platzer, E. G., Bellier, A. & Aroian, R. V. Discovery of a highly synergistic anthelmintic combination that shows mutual hypersusceptibility. Proc. Natl. Acad. Sci. USA 107, 5955–5960 (2010).
Burns, A. R. et al. A predictive model for drug bioaccumulation and bioactivity in Caenorhabditis elegans. Nat. Chem. Biol. 6, 549–557 (2010).
Ortiz de Montellano, P. R. Cytochrome P450-activated prodrugs. Future Med. Chem. 5, 213–228 (2013).
Leung, M. C. K., Goldstone, J. V., Boyd, W. A., Freedman, J. H. & Meyer, J. N. Caenorhabditis elegans generates biologically relevant levels of genotoxic metabolites from aflatoxin B1 but not benzo[a]pyrene in vivo. Toxicol. Sci. 118, 444–453 (2010).
Harlow, P. H., Perry, S. J., Stevens, A. J. & Flemming, A. J. Comparative metabolism of xenobiotic chemicals by cytochrome P450s in the nematode Caenorhabditis elegans. Sci Rep. 8, 13333 (2018).
Porter, T. D. New insights into the role of cytochrome P450 reductase (POR) in microsomal redox biology. Acta Pharm. Sin. B 2, 102–106 (2012).
Kalgutkar, A. S. et al. Reactive metabolite trapping studies on imidazo- and 2-methylimidazo[2,1-b]thiazole-based inverse agonists of the ghrelin receptor. Drug Metab. Dispos. 7, 1375–1388 (2013).
Ryan, E. et al. Evidence for the in vitro bioactivation of aminopyrazole derivatives: trapping reactive aminopyrazole intermediates using glutathione ethyl ester in human liver microsomes. Chem. Res. Toxicol. 28, 1747–1752 (2015).
Cooper, A. J. L. & Hanigan, M. H. Metabolism of glutathione S-conjugates: multiple pathways. Compr. Toxicol. https://doi.org/10.1016/B978-0-12-801238-3.01973-5 (2018).
Ferguson, G. D. & Bridge, W. J. The glutathione system and the related thiol network in Caenorhabditis elegans. Redox Biol. 24, 101171 (2019).
Blum, R., Meyer, K. C., Wünschmann, J., Lendzian, K. J. & Grill, E. Cytosolic action of phytochelatin synthase. Plant Physiol. 153, 159–169 (2010).
Essig, Y. J., Webb, S. M. & Stürzenbaum, S. R. Deletion of phytochelatin synthase modulates the metal accumulation pattern of cadmium exposed C. elegans. Int. J. Mol. Sci. 17, 257 (2016).
Giustarini, D., Milzani, A., Dalle-Donne, I., Tsikas, D. & Rossi, R. N-Acetylcysteine ethyl ester (NACET): a novel lipophilic cell-permeable cysteine derivative with an unusual pharmacokinetic feature and remarkable antioxidant potential. Biochem. Pharmacol. 84, 1522–1533 (2012).
Trudgill, D. L. & Blok, V. C. Apomictic, polyphagus root-knot nematodes: exceptionally successful and damaging biotrophic root pathogens. Annu. Rev. Phytopathol. 39, 53–77 (2001).
Oka, Y. From old-generation to next-generation nematicides. Agronomy 10, 1387 (2020).
Devguard 500SC Label,https://za.uplonline.com/download_links/awyXPTiWs8OT3dGCErYdw3nnA3nLnwxpCpUn7jzb.pdf (deVGen/UPL).
Asif, M., Rehman, B., Parihar, K., Ganai, M. A. & Siddiqui, M. A. Effect of various physico-chemical factors on the incidence of root knot nematode Meloidogyne spp. infesting tomato in District Aligarh (Uttar Pradesh) India. J. Plant Sci. 10, 234–243 (2015).
Barker, K. R., Schmitt, D. P. & Imbriani, J. L. in An Advanced Treatise on Meloidogyne Volume II: Methodology (eds Barker, K. R. et al.) 135–148 (North Carolina State University Department of Plant Pathology and the United States Agency for International Development, 1985).
Marion, M. J., Hantz, O. & Durantel, D. in Hepatocytes: Methods and Protocols (ed. Maurel, P.) 261–272 (Humana Press, 2010).
Rocher, F. et al. Salicylic acid transport in Ricinus communis involves a pH-dependent carrier system in addition to diffusion. Plant Physiol. 150, 2081–2091 (2009).
Wram, C. L., Hesse, C. N. & Zasada, I. A. Transcriptional response of Meloidogyne incognita to non-fumigant nematicides. Sci Rep. 12, 9814 (2022).
Dara, S. K. The new integrated pest management paradigm for the modern age. J. Integr. Pest. Manag. 10, 12 (2019).
Cardarelli, M., Woo, S. L., Rouphael, Y. & Colla, G. Seed treatments with microorganisms can have a biostimulant effect by influencing germination and seedling growth of crops. Plants 11, 259 (2022).
Migunova, V. D. & Sasanelli, N. Bacteria as biocontrol tool against phytoparasitic nematodes. Plants 10, 389 (2021).
Slomczynska, U. et al. in Discovery and Synthesis of Crop Protection Products (eds. Maienfisch, P. & Stevenson, T.) 129–147 (American Chemical Society, 2015).
Jensen, J. P., Kalwa, U., Pandey, S. & Tylka, G. L. Avicta and clariva affect the biology of the soybean cyst nematode, Heterodera glycines. Plant Dis. 102, 2480–2486 (2018).
Ahmed, S., Zhou, Z., Zhou, J. & Chen, S.-Q. Pharmacogenomics of drug metabolizing enzymes and transporters: relevance to precision medicine. Genomics Proteomics Bioinformatics 14, 298–313 (2016).
Nixon, S. A. et al. Where are all the anthelmintics? Challenges and opportunities on the path to new anthelmintics. Int. J. Parasitol. Drugs Drug Resist. 14, 8–16 (2020).
Lewis, J. A. & Fleming, J. T. in Caenorhabditis elegans: Modern Biological Analysis of an Organism (eds Epstein, H. F. & Shakes, D. C.) 3–29 (Academic Press, 1995).
Burns, A. R. et al. High-throughput screening of small molecules for bioactivity and target identification in Caenorhabditis elegans. Nat. Protoc. 1, 1906–1914 (2006).
Demeler, J., Kuttler, U. & von Samson-Himmelstjerna, G. Veterinary parasitology adaptation and evaluation of three different in vitro tests for the detection of resistance to anthelmintics in gastro intestinal nematodes of cattle. Vet. Parasitol. 170, 61–70 (2010).
Bartley, D. J. et al. A survey of anthelmintic resistant nematode parasites in Scottish sheep flocks. Vet. Parasitol. 117, 61–71 (2003).
Jindapunnapat, K., Reetz, N. D., Macdonald, M. H., Bhagavathy, G. & Meyer, S. L. F. Activity of vetiver extracts and essential oil against Meloidogyne incognita. J. Nematol. 50, 147–162 (2018).
Wram, C. L. & Zasada, I. A. Short-term effects of sublethal doses of nematicides on Meloidogyne incognita. Phytopathology 109, 1605–1613 (2019).
Meyer, S. L. F., Chauhan, K. R. & MacDonald, M. H. Evaluation of roselle (Hibiscus sabdariffa) leaf and pomegranate (Punica granatum) fruit rind for activity against Meloidogyne incognita. Nematropica 46, 85–96 (2016).
Meyer, S. L. F. et al. Plantago lanceolata and Plantago rugelii extracts are toxic to Meloidogyne incognita but not to certain microbes. J. Nematol. 38, 333–338 (2006).
Sherman, F. Getting started with yeast. Methods Enzymol. 350, 3–41 (2002).
Howe, K. L. et al. WormBase ParaSite—a comprehensive resource for helminth genomics. Mol. Biochem. Parasitol. 215, 2–10 (2016).
Blanc-Mathieu, R. et al. Hybridization and polyploidy enable genomic plasticity without sex in the most devastating plant-parasitic nematodes. PLoS Genet. 13, e1006777 (2017).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
WormBase release WS283, http://www.wormbase.org (Alliance of Genome Resources, accessed 16 December 2021).
Wram, C. L. & Zasada, I. Differential response of Meloidogyne, Pratylenchus, Globodera, and Xiphinema species to the nematicide fluazaindolizine. Phytopathology. 110, 2003–2009 (2020).
Chundawat, T. S., Kumari, P., Sharma, N. & Bhagat, S. Strategic synthesis and in vitro antimicrobial evaluation of novel difluoromethylated 1-(1,3-diphenyl-1H-pyrazol-4-yl)-3,3-difluoro-1,3-dihydro-indol-2-ones. Med. Chem. Res. 25, 2335–2348 (2016).
Pyl, T., Giebelmann, R. & Beyer, H. Über bicyclische heterocyclen mit gemeinsamem stickstoffatom, I. Zur kenntnis der imidazo[2.1-b]thiazole. Justus Liebigs Ann. Chem. 643, 145–153 (1961).
Huang, G. et al. Engineering broad root-knot resistance in transgenic plants by RNAi silencing of a conserved and essential root-knot nematode parasitism gene. Proc. Natl Acad. Sci. USA 103, 14302–14306 (2006).
Mota, F. C. et al. New sources of resistance to Meloidogyne incognita race 3 in wild cotton accessions and histological characterization of the defence mechanisms. Plant Pathol. 62, 1173–1183 (2013).
Basso, M. F. et al. MiDaf16-like and MiSkn1-like gene families are reliable targets to develop biotechnological tools for the control and management of Meloidogyne incognita. Sci Rep. 10, 6991 (2020).
Hajihassani, A., Rutter, W. B. & Luo, X. Resistant pepper carrying N, Me1, and Me3 have different effects on penetration and reproduction of four major Meloidogyne species. J. Nematol. 51, 1–9 (2019).
de Souza, J. D. A. et al. Knocking-down Meloidogyne incognita proteases by plant-delivered dsRNA has negative pleiotropic effect on nematode vigor. PLoS ONE 8, e85364 (2013).
Burns, A. R. and Roy, P. J. Python and R scripts for the generation of dose–response heatmaps and heatmaps of HPLC chromatograms. Zenodo https://doi.org/10.5281/zenodo.7731172 (2023).





More News
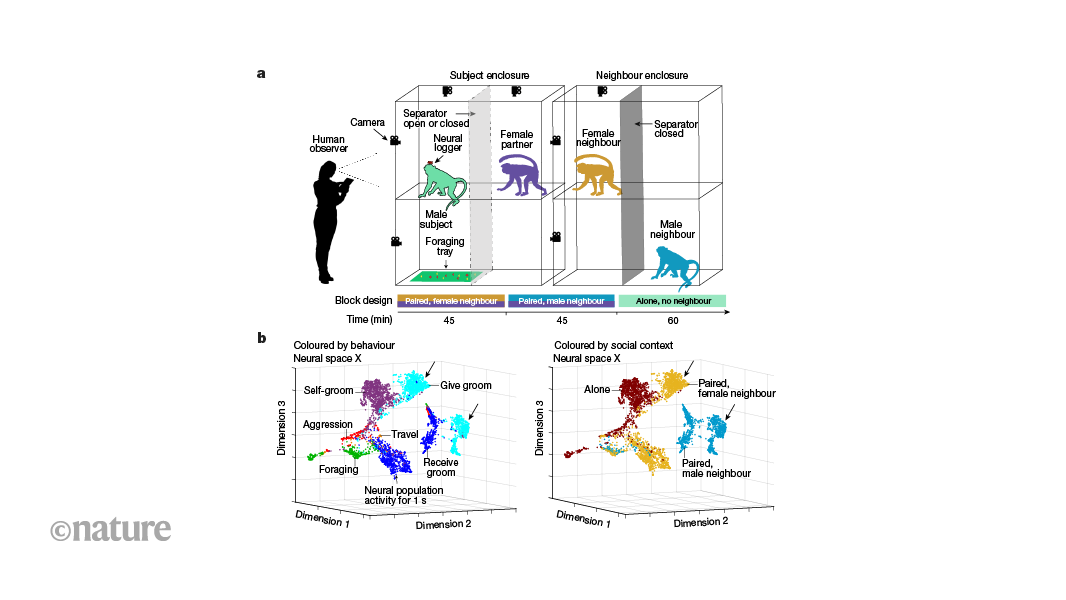
Monkey business: primates’ social life tracked with wireless neuronal recording
Daily briefing: Carrion crows have counting skills seen only in people
Researcher parents are paying a high price for conference travel — here’s how to fix it